APL SURP Python course - Notebook 4 (completed version)#
If statements, for loops, xarray, and netCDF files
Created for the University of Washington Applied Physics Laboratory’s Summer Undergraduate Research Program (SURP) 2025.
For additional resources on Python basics, you can consult the following resources on the APL-SURP Python course website:
Tutorials on Python fundamentals: https://uw-apl-surp.github.io/aplsurp-python/overview.html
Complementary lessons on specific Python topics: https://uw-apl-surp.github.io/aplsurp-python/complementary_lessons.html
import numpy as np # NumPy is an array and math library
import matplotlib.pyplot as plt # Matplotlib is a visualization (plotting) library
import pandas as pd # Pandas lets us work with spreadsheet (.csv) data
from datetime import datetime, timedelta # Datetime helps us work with dates and times
import xarray as xr # (NEW!) xarray handles multidimensional labeled data
Part 1: if statements and for loops#
if statements#
We can use logical operations to create conditional actions using the if statement.
If the boolean condition evaluates to True, then the lines inside the first block are executed.
Important: note the colons (:) and how the lines below each condition are indented using a Tab or two spaces on your keyboard (you can use a different number of spaces, but they have to be the same within a block).
if <CONDITION #1>:
<ACTION>
<ACTION>
etc.
To create multiple paths for the code to follow, you can use an if-else statement.
If the first condition evaluates to True, then the lines inside the first block are executed.
If the first condition evaluates to False, then Python tests the second elif statement.
If all of the if and elif statements are False, then Python will finally execute the else statement.
if <CONDITION #1>:
<ACTION>
<ACTION>
etc.
elif <CONDITION #2>: (optional)
<ACTION>
elif <CONDITION #3>: (optional)
<ACTION>
else: (optional)
<ACTION>
First, a quick review of logical operations and booleans. What will these four print statements tell us?
current_temp = 70.0 # 70°F
print(current_temp >= 50)
print(current_temp >= 60)
print(current_temp >= 70)
print(current_temp >= 80)
True
True
True
False
Now, try changing the value of rain_chance below and running the code. Do you understand the control flow?
What range of values for rain_chance will trigger the else statement?
rain_chance = 5 # i.e., a 5% chance of rain
if rain_chance >= 50:
print('Ugh... I better bring an umbrella.')
elif rain_chance == 0:
print('I definitely will not need an umbrella.')
elif rain_chance <= 20:
print('I should be okay without an umbrella.')
else:
print('I am not sure what to do.')
I should be okay without an umbrella.
for loops#
Sometimes, we might want to perform an action multiple times, with small modifications. Coding makes this possible using loops!
A for loop has the following syntax:
for <ELEMENT> in <ITERABLE>:
<ACTION>
<ACTION>
etc.
Here, <ITERABLE> can be a list, array, string, or other collection of elements. You can give <ELEMENT> any unique variable name, and that variable will typically only be used inside the loop.
Run and consider the following examples:
countdown = [4,3,2,1]
for item in countdown:
print(item)
4
3
2
1
for character in 'floats':
print(character)
f
l
o
a
t
s
for num in np.arange(0,7,2):
print(num)
0
2
4
6
# These are the monthly average temperatures for Seattle from before
seattle_temps = np.array([40.0,40.6,44.2,48.4,54.9,60.2,66.2,66.7,60.5,52.0,44.5,39.6])
# This loop counts the number of months in which temps are warmer than 60°F:
warm_months = 0
for month in np.arange(0,12):
this_month_temp = seattle_temps[month]
if this_month_temp > 60.0:
warm_months = warm_months + 1
print(f'On average, there are {warm_months} warm months in Seattle!')
On average, there are 4 warm months in Seattle!
You already learned how to calculate the sum of an array of numbers using np.sum().
Now, try to calculate the average Seattle annual temperature by writing a for loop below. There are at least two different ways to write the loop!
# You can reuse the variable "seattle_temps" defined earlier
# Write your code below:
# Option 1:
sum_temp = 0
for temp in seattle_temps:
sum_temp += temp
# Option 2:
sum_temp = 0
for index in np.arange(len(seattle_temps)):
sum_temp += seattle_temps[index]
# Finally, print the average temperature by adding it to the print() statement:
print(f'The average annual temperature is: {sum_temp / len(seattle_temps):.2f} °F')
# Here we give equal weight to every month, so this isn't fully accurate
The average annual temperature is: 51.48 °F
Note: Numpy arrays and Pandas DataFrames provide capabilities that often make the use of if and for statements unnecessary. Logical operations can generate boolean indices applied over arrays, and functions can be applied over all elements in an array or column.
Part 2: xarray and netCDF files#
Intro to netCDF files#
We’ve been working with NumPy arrays, and we know that they have axes. A 3-D array, for instance, has 3 axes. NumPy calls them axes 0, 1, and 2, but you could also think of them as x, y, and z, or longitude, latitude, and depth. And we know from indexing that we can identify a value by its coordinates along those axes.
Image source: digitalearthafrica.org
You will often find multidimensional data is from climate or weather models, satellite products, and anything else that can be shown on a map.
In these data sets, Earth is often divided into grid cells in order to simplify the challenge of mapping parameters like ocean temperature or cloud cover all around the world.
As the image below shows, those grid cells often extend in both the horizontal dimension (latitude and longitude) as well as the vertical dimension (height, or depth).
Image source: Caltech
xarray is an important community-developed package that lets us work with gridded data.
In science, gridded data is generally provided using netCDF files, which will usually (but not always) have the file extension .nc.
Because netCDF files are compressed, you can’t simply open them on your computer in TextEdit or Notepad or Excel or Google Sheets. Most of the time, we have to write code to look inside them.
Image source: Matthew Rocklin
The image above is what you might see when you open a netCDF file in xarray. The file may contain multiple variables: in this example, temperature, pressure, elevation, and land_cover.
Those variables will be aligned along a few dimensions — in this case, latitude, longitude, and time.
But not every variable will use all the dimensions. Here, temperature is 3-D, while elevation is only 2-D, because the land elevation doesn’t change much over time (at least over human time scales!).
The dimensions themselves will have coordinates, which are the indices with numbers that let you locate a certain data value along a dimension. A set of coordinates might be: latitude = 70°N, longitude = 130°W (which is the same as -130°E), and time = datetime(2025,7,24).
Lastly, netCDF files have attributes, which are just simple pieces of text (or “metadata”) that may describe how the data set was created, the version of the data set, the units of variables, and other useful info.
Intro to xarray#
We’ve already imported xarray at the top using:
import xarray as xr
Now, let’s download a netCDF file, which is located on Google Drive here.
This is satellite ocean color data, which gives an estimation of chlorophyll concentration—itself a measure of phytoplankton in the ocean surface. It is from NOAA’s VIIRS sensor, provided at a fine resolution of 750 m (0.75 km) over 8-day composites. The data was obtained from this NOAA ERDDAP server.
We load the netCDF file into an xarray Dataset object using the xr.open_dataset() function, which takes the filepath (including filename) as its argument. You can find its documentation here.
Note that there is also a similar function called xr.open_mfdataset() that opens and combines multiple netCDF files.
# Load the netCDF file and display its contents
ds = xr.open_dataset('/content/erdVHNchla8day_5d67_fb51_b559_U1753127354803.nc')
display(ds) # In a notebook, you can also just type "ds" without display()
<xarray.Dataset> Size: 14MB
Dimensions: (time: 111, altitude: 1, latitude: 268, longitude: 121)
Coordinates:
* time (time) datetime64[ns] 888B 2025-03-01 2025-03-02 ... 2025-07-16
* altitude (altitude) float64 8B 0.0
* latitude (latitude) float64 2kB 49.0 48.99 48.99 ... 47.01 47.01 47.0
* longitude (longitude) float64 968B -123.1 -123.1 -123.1 ... -122.2 -122.2
Data variables:
chla (time, altitude, latitude, longitude) float32 14MB ...
Attributes: (12/47)
cdm_data_type: Grid
contact: erd.data@noaa.gov
Conventions: CF-1.6, COARDS, ACDD-1.3
creation_date: Mon Jul 21 03:13:41 2025
creator_email: erd.data@noaa.gov
creator_name: NOAA NMFS SWFSC ERD
... ...
standard_name_vocabulary: CF Standard Name Table v70
summary: NOAA CoastWatch distributes L3 level chlor...
time_coverage_end: 2025-07-16T00:00:00Z
time_coverage_start: 2025-03-01T00:00:00Z
title: Chlorophyll a, North Pacific, NOAA VIIRS, ...
Westernmost_Easting: -123.10125We see that the file has 4 dimensions, 4 coordinates, 1 variable, and 47 attributes.
One of the dimensions only has a single coordinate value. Which is it?
Use the interactive view to explore the data.
What is the full name of the chla variable, and what units is it in?
What is the range of longitudes in the data set, expressed as degrees W (°W)?
Note that latitude values are reversed relative to the usual expectation in netCDF files that coordinate values are in increasing order.
To extract a single variable or coordinate, we generally use bracket syntax:
dataset_name['variable_name_as_a_string']
Another option is to use dot-syntax:
dataset_name.variable_name
Either way, this will give us an xarray DataArray object, which is basically a wrapper around a NumPy array:
# Example: extract just the time coordinate
ds['time']
# Note that this gives the same result:
# ds.time
<xarray.DataArray 'time' (time: 111)> Size: 888B
array(['2025-03-01T00:00:00.000000000', '2025-03-02T00:00:00.000000000',
'2025-03-03T00:00:00.000000000', '2025-03-04T00:00:00.000000000',
'2025-03-05T00:00:00.000000000', '2025-03-06T00:00:00.000000000',
'2025-03-07T00:00:00.000000000', '2025-03-08T00:00:00.000000000',
'2025-03-09T00:00:00.000000000', '2025-03-10T00:00:00.000000000',
'2025-03-13T00:00:00.000000000', '2025-03-14T00:00:00.000000000',
'2025-03-15T00:00:00.000000000', '2025-03-16T00:00:00.000000000',
'2025-03-17T00:00:00.000000000', '2025-03-18T00:00:00.000000000',
'2025-03-19T00:00:00.000000000', '2025-03-20T00:00:00.000000000',
'2025-03-21T00:00:00.000000000', '2025-03-22T00:00:00.000000000',
'2025-03-23T00:00:00.000000000', '2025-03-27T00:00:00.000000000',
'2025-03-28T00:00:00.000000000', '2025-03-29T00:00:00.000000000',
'2025-03-30T00:00:00.000000000', '2025-03-31T00:00:00.000000000',
'2025-04-01T00:00:00.000000000', '2025-04-02T00:00:00.000000000',
'2025-04-03T00:00:00.000000000', '2025-04-04T00:00:00.000000000',
'2025-04-05T00:00:00.000000000', '2025-04-06T00:00:00.000000000',
'2025-04-07T00:00:00.000000000', '2025-04-08T00:00:00.000000000',
'2025-04-09T00:00:00.000000000', '2025-04-10T00:00:00.000000000',
'2025-04-11T00:00:00.000000000', '2025-04-12T00:00:00.000000000',
'2025-04-13T00:00:00.000000000', '2025-04-14T00:00:00.000000000',
'2025-04-15T00:00:00.000000000', '2025-04-16T00:00:00.000000000',
'2025-04-17T00:00:00.000000000', '2025-04-18T00:00:00.000000000',
'2025-04-19T00:00:00.000000000', '2025-04-20T00:00:00.000000000',
'2025-04-21T00:00:00.000000000', '2025-04-22T00:00:00.000000000',
'2025-04-23T00:00:00.000000000', '2025-04-24T00:00:00.000000000',
'2025-04-25T00:00:00.000000000', '2025-04-26T00:00:00.000000000',
'2025-04-27T00:00:00.000000000', '2025-04-28T00:00:00.000000000',
'2025-04-29T00:00:00.000000000', '2025-04-30T00:00:00.000000000',
'2025-05-01T00:00:00.000000000', '2025-05-02T00:00:00.000000000',
'2025-05-03T00:00:00.000000000', '2025-05-04T00:00:00.000000000',
'2025-05-05T00:00:00.000000000', '2025-05-06T00:00:00.000000000',
'2025-05-07T00:00:00.000000000', '2025-05-08T00:00:00.000000000',
'2025-05-09T00:00:00.000000000', '2025-05-10T00:00:00.000000000',
'2025-05-11T00:00:00.000000000', '2025-05-12T00:00:00.000000000',
'2025-05-13T00:00:00.000000000', '2025-05-14T00:00:00.000000000',
'2025-05-15T00:00:00.000000000', '2025-05-16T00:00:00.000000000',
'2025-05-17T00:00:00.000000000', '2025-05-18T00:00:00.000000000',
'2025-05-19T00:00:00.000000000', '2025-05-20T00:00:00.000000000',
'2025-05-21T00:00:00.000000000', '2025-05-22T00:00:00.000000000',
'2025-05-23T00:00:00.000000000', '2025-05-26T00:00:00.000000000',
'2025-05-27T00:00:00.000000000', '2025-05-28T00:00:00.000000000',
'2025-05-29T00:00:00.000000000', '2025-05-30T00:00:00.000000000',
'2025-05-31T00:00:00.000000000', '2025-06-01T00:00:00.000000000',
'2025-06-02T00:00:00.000000000', '2025-06-03T00:00:00.000000000',
'2025-06-04T00:00:00.000000000', '2025-06-05T00:00:00.000000000',
'2025-06-06T00:00:00.000000000', '2025-06-07T00:00:00.000000000',
'2025-06-08T00:00:00.000000000', '2025-06-09T00:00:00.000000000',
'2025-06-10T00:00:00.000000000', '2025-06-11T00:00:00.000000000',
'2025-06-12T00:00:00.000000000', '2025-06-21T00:00:00.000000000',
'2025-06-22T00:00:00.000000000', '2025-07-05T00:00:00.000000000',
'2025-07-06T00:00:00.000000000', '2025-07-07T00:00:00.000000000',
'2025-07-08T00:00:00.000000000', '2025-07-09T00:00:00.000000000',
'2025-07-10T00:00:00.000000000', '2025-07-11T00:00:00.000000000',
'2025-07-12T00:00:00.000000000', '2025-07-13T00:00:00.000000000',
'2025-07-14T00:00:00.000000000', '2025-07-15T00:00:00.000000000',
'2025-07-16T00:00:00.000000000'], dtype='datetime64[ns]')
Coordinates:
* time (time) datetime64[ns] 888B 2025-03-01 2025-03-02 ... 2025-07-16
Attributes:
_CoordinateAxisType: Time
actual_range: [1.7407872e+09 1.7526240e+09]
axis: T
ioos_category: Time
long_name: Centered Time
standard_name: time
time_origin: 01-JAN-1970 00:00:00We can retrieve the underlying NumPy array using the following .values or .data syntax. In this case, we get a NumPy array filled with datetime objects:
# Convert the data from xarray into NumPy
ds['time'].values
# Note that this gives the same result:
# ds['time'].data
array(['2025-03-01T00:00:00.000000000', '2025-03-02T00:00:00.000000000',
'2025-03-03T00:00:00.000000000', '2025-03-04T00:00:00.000000000',
'2025-03-05T00:00:00.000000000', '2025-03-06T00:00:00.000000000',
'2025-03-07T00:00:00.000000000', '2025-03-08T00:00:00.000000000',
'2025-03-09T00:00:00.000000000', '2025-03-10T00:00:00.000000000',
'2025-03-13T00:00:00.000000000', '2025-03-14T00:00:00.000000000',
'2025-03-15T00:00:00.000000000', '2025-03-16T00:00:00.000000000',
'2025-03-17T00:00:00.000000000', '2025-03-18T00:00:00.000000000',
'2025-03-19T00:00:00.000000000', '2025-03-20T00:00:00.000000000',
'2025-03-21T00:00:00.000000000', '2025-03-22T00:00:00.000000000',
'2025-03-23T00:00:00.000000000', '2025-03-27T00:00:00.000000000',
'2025-03-28T00:00:00.000000000', '2025-03-29T00:00:00.000000000',
'2025-03-30T00:00:00.000000000', '2025-03-31T00:00:00.000000000',
'2025-04-01T00:00:00.000000000', '2025-04-02T00:00:00.000000000',
'2025-04-03T00:00:00.000000000', '2025-04-04T00:00:00.000000000',
'2025-04-05T00:00:00.000000000', '2025-04-06T00:00:00.000000000',
'2025-04-07T00:00:00.000000000', '2025-04-08T00:00:00.000000000',
'2025-04-09T00:00:00.000000000', '2025-04-10T00:00:00.000000000',
'2025-04-11T00:00:00.000000000', '2025-04-12T00:00:00.000000000',
'2025-04-13T00:00:00.000000000', '2025-04-14T00:00:00.000000000',
'2025-04-15T00:00:00.000000000', '2025-04-16T00:00:00.000000000',
'2025-04-17T00:00:00.000000000', '2025-04-18T00:00:00.000000000',
'2025-04-19T00:00:00.000000000', '2025-04-20T00:00:00.000000000',
'2025-04-21T00:00:00.000000000', '2025-04-22T00:00:00.000000000',
'2025-04-23T00:00:00.000000000', '2025-04-24T00:00:00.000000000',
'2025-04-25T00:00:00.000000000', '2025-04-26T00:00:00.000000000',
'2025-04-27T00:00:00.000000000', '2025-04-28T00:00:00.000000000',
'2025-04-29T00:00:00.000000000', '2025-04-30T00:00:00.000000000',
'2025-05-01T00:00:00.000000000', '2025-05-02T00:00:00.000000000',
'2025-05-03T00:00:00.000000000', '2025-05-04T00:00:00.000000000',
'2025-05-05T00:00:00.000000000', '2025-05-06T00:00:00.000000000',
'2025-05-07T00:00:00.000000000', '2025-05-08T00:00:00.000000000',
'2025-05-09T00:00:00.000000000', '2025-05-10T00:00:00.000000000',
'2025-05-11T00:00:00.000000000', '2025-05-12T00:00:00.000000000',
'2025-05-13T00:00:00.000000000', '2025-05-14T00:00:00.000000000',
'2025-05-15T00:00:00.000000000', '2025-05-16T00:00:00.000000000',
'2025-05-17T00:00:00.000000000', '2025-05-18T00:00:00.000000000',
'2025-05-19T00:00:00.000000000', '2025-05-20T00:00:00.000000000',
'2025-05-21T00:00:00.000000000', '2025-05-22T00:00:00.000000000',
'2025-05-23T00:00:00.000000000', '2025-05-26T00:00:00.000000000',
'2025-05-27T00:00:00.000000000', '2025-05-28T00:00:00.000000000',
'2025-05-29T00:00:00.000000000', '2025-05-30T00:00:00.000000000',
'2025-05-31T00:00:00.000000000', '2025-06-01T00:00:00.000000000',
'2025-06-02T00:00:00.000000000', '2025-06-03T00:00:00.000000000',
'2025-06-04T00:00:00.000000000', '2025-06-05T00:00:00.000000000',
'2025-06-06T00:00:00.000000000', '2025-06-07T00:00:00.000000000',
'2025-06-08T00:00:00.000000000', '2025-06-09T00:00:00.000000000',
'2025-06-10T00:00:00.000000000', '2025-06-11T00:00:00.000000000',
'2025-06-12T00:00:00.000000000', '2025-06-21T00:00:00.000000000',
'2025-06-22T00:00:00.000000000', '2025-07-05T00:00:00.000000000',
'2025-07-06T00:00:00.000000000', '2025-07-07T00:00:00.000000000',
'2025-07-08T00:00:00.000000000', '2025-07-09T00:00:00.000000000',
'2025-07-10T00:00:00.000000000', '2025-07-11T00:00:00.000000000',
'2025-07-12T00:00:00.000000000', '2025-07-13T00:00:00.000000000',
'2025-07-14T00:00:00.000000000', '2025-07-15T00:00:00.000000000',
'2025-07-16T00:00:00.000000000'], dtype='datetime64[ns]')
Can you use code to print the time range of the data — in other words, the start and end dates?
# Write your code here:
print(ds['time'].values.min(),ds['time'].values.max())
2025-03-01T00:00:00.000000000 2025-07-16T00:00:00.000000000
Selecting data in xarray#
To select one or more values inside an xarray variable, we use .isel() to select by integer index or .sel() to select by coordinate value:
For example, if we wanted to select the most recent time value of chlorophyll, we’d use selection by index:
# Select chla at the most recent time:
ds['chla'].isel(time=-1)
<xarray.DataArray 'chla' (altitude: 1, latitude: 268, longitude: 121)> Size: 130kB
[32428 values with dtype=float32]
Coordinates:
time datetime64[ns] 8B 2025-07-16
* altitude (altitude) float64 8B 0.0
* latitude (latitude) float64 2kB 49.0 48.99 48.99 ... 47.01 47.01 47.0
* longitude (longitude) float64 968B -123.1 -123.1 -123.1 ... -122.2 -122.2
Attributes:
colorBarMaximum: 30.0
colorBarMinimum: 0.03
colorBarScale: Log
coordsys: geographic
ioos_category: Ocean Color
long_name: Chlorophyll a
standard_name: mass_concentration_chlorophyll_concentration_in_sea_water
units: mg m^-3
valid_max: 100.1
valid_min: 0.001If we knew we wanted to find the chlorophyll on April 1, 2025 at a latitude of 48.00375°N and a longitude of 122.35125°W, we’d write:
# Select chla at the specific time/lat/lon coordinates:
ds['chla'].sel(time=datetime(2025,4,1),latitude=48.00375,longitude=-122.35125)
# Note the longitude values are negative for degrees W (°W), since the units are °E
<xarray.DataArray 'chla' (altitude: 1)> Size: 4B
[1 values with dtype=float32]
Coordinates:
time datetime64[ns] 8B 2025-04-01
* altitude (altitude) float64 8B 0.0
latitude float64 8B 48.0
longitude float64 8B -122.4
Attributes:
colorBarMaximum: 30.0
colorBarMinimum: 0.03
colorBarScale: Log
coordsys: geographic
ioos_category: Ocean Color
long_name: Chlorophyll a
standard_name: mass_concentration_chlorophyll_concentration_in_sea_water
units: mg m^-3
valid_max: 100.1
valid_min: 0.001Sometimes we need to add an .item() to pull a single data value out from xarray so we can see it:
# Select chla at the specific time/lat/lon coordinates:
ds['chla'].sel(time=datetime(2025,4,1),latitude=48.00375,longitude=-122.35125).item()
2.5553884506225586
Plotting data from xarray#
Just like Pandas, xarray comes with built-in plotting functions that are similar to those in Matplotlib.
You can usually add .plot() after an xarray 1-D or 2-D variable to create a simple plot.
For example, let’s plot the chlorophyll on the most recent day of data:
# Create a quick plot
ds['chla'].isel(time=-1).plot()
<matplotlib.collections.QuadMesh at 0x7d7c7ec5ef50>
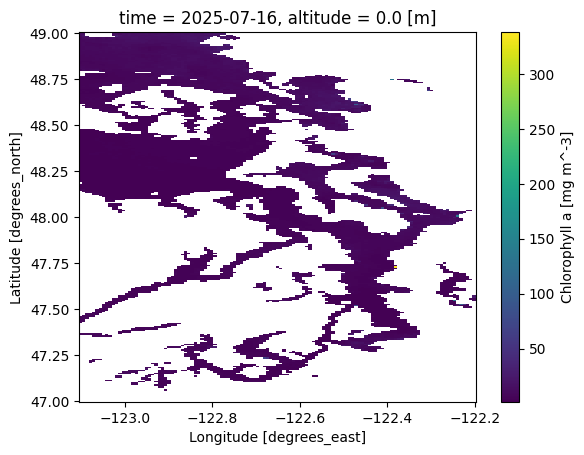
Notice that the color range seems to be too big. Most of the data has much lower chlorophyll values!
Let’s set the color range to have a maximum value of 20 mg/m^3 using the vmax argument.
Can you find Seattle on this map of Puget Sound?
# Specify the maximum color range
ds['chla'].isel(time=-1).plot(vmax=20.0)
<matplotlib.collections.QuadMesh at 0x7d7c7e9eecd0>
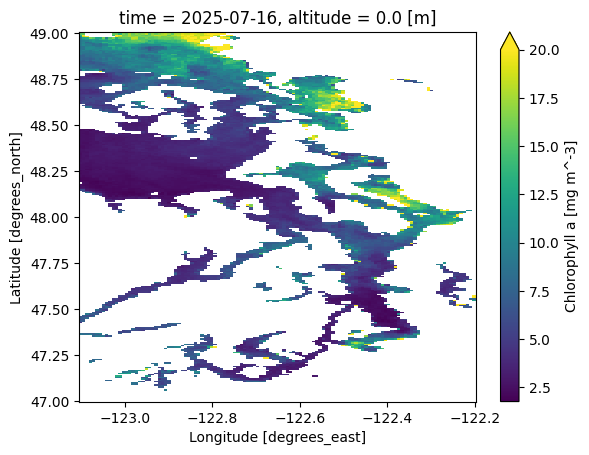
Just like NumPy arrays or Pandas DataFrames and Series, you can do math using xarray objects!
The beauty of xarray is that it will align coordinates to make sure the right values are being added, subtracted, multiplied, or divided.
Can you calculate how chlorophyll has changed between the first day of data and the most recent day of data, then create a plot of the change?
# Write your code here:
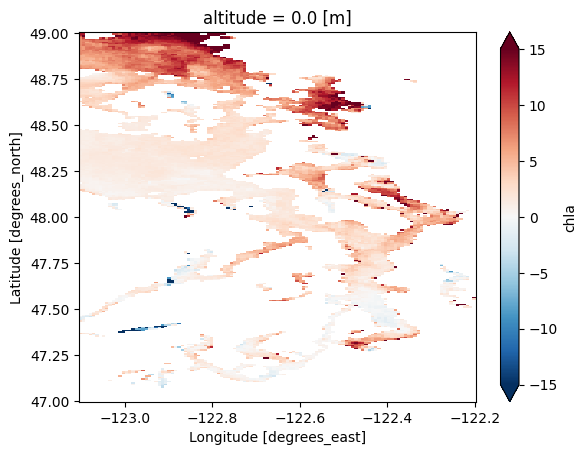
chl_diff = ds['chla'].isel(time=-1) - ds['chla'].isel(time=0)
chl_diff.plot(robust=True)
# Note that adding "robust=True" inside .plot() removes outlier points outside the
# 2nd and 98th percentiles, making it easier to visualize the data.
# We could also write:
# chl_diff.plot(vmin=-15,vmax=15,cmap='RdBu_r')
<matplotlib.collections.QuadMesh at 0x7d7c7a1a5150>
Manipulating xarray data#
Similar to Pandas, you can apply NumPy functions like .mean() to an xarray object.
For example, we can calculate the average across the longitude dimension—in other words, across all longitudes. That leaves time, latitude, and altitude as the remaining dimensions:
data.mean(dim='longitude')
Let’s calculate the average chlorophyll over time within the central Puget Sound region. Here we’ll specify two dimensions inside .mean(): latitude and longitude.
Notice how these operations form a chain:
Variable selection, then
Coordinate selection/slicing, then
Dimension reduction (averaging), then
Finally, plotting!
ds['chla'].sel(longitude=slice(-122.55,-122.3),
latitude=slice(48,47.50)).mean(dim=['longitude',
'latitude']).plot()
[<matplotlib.lines.Line2D at 0x7d7c79d8f590>]
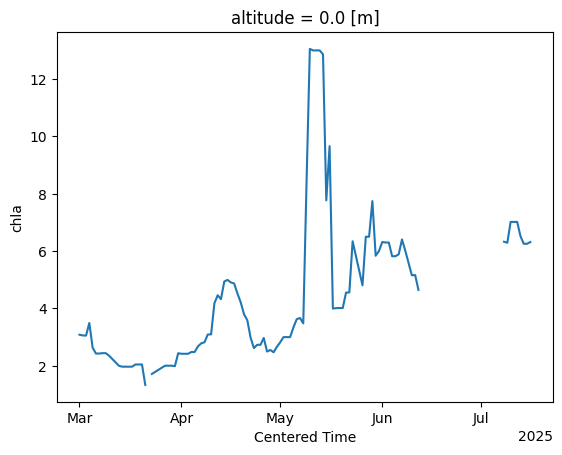
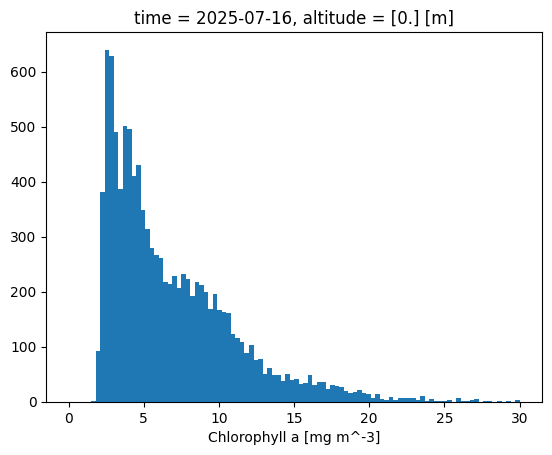
Another neat quick plotting option is the histogram, .plot.hist(), which shows the distribution of chlorophyll values within specified bins:
ds['chla'].isel(time=-1).plot.hist(bins=100,range=[0,30]);