APL SURP Python course - Notebook 3 (completed version)#
Line and scatter plots, depth profiles, timeseries data, logical operations, if statements and for loops and more
Created for the University of Washington Applied Physics Laboratory’s Summer Undergraduate Research Program (SURP) 2025.
For additional resources on Python basics, you can consult the following resources on the APL-SURP Python course website:
Tutorials on Python fundamentals: https://uw-apl-surp.github.io/aplsurp-python/overview.html
Complementary lessons on specific Python topics: https://uw-apl-surp.github.io/aplsurp-python/complementary_lessons.html
import numpy as np # NumPy is an array and math library
import matplotlib.pyplot as plt # Matplotlib is a visualization (plotting) library
import pandas as pd # Pandas lets us work with spreadsheet (.csv) data
from datetime import datetime, timedelta # Datetime helps us work with dates and times
Part 1: Line and scatter plots#
It’s time for us to start creating visualizations of data, called plots.
At the top of this page, we imported the package Matplotlib using:
import matplotlib.pyplot as plt
Creating a line plot is simple. We use the Matplotlib function plt.plot(). The basic form of the function is:
plt.plot(X, Y, <FORMAT_ARGUMENTS>...)
Here, X and Y should be 1-D lists or arrays of data. The options for <FORMAT_ARGUMENTS> can be found on Matplotlib’s documentation webpage.
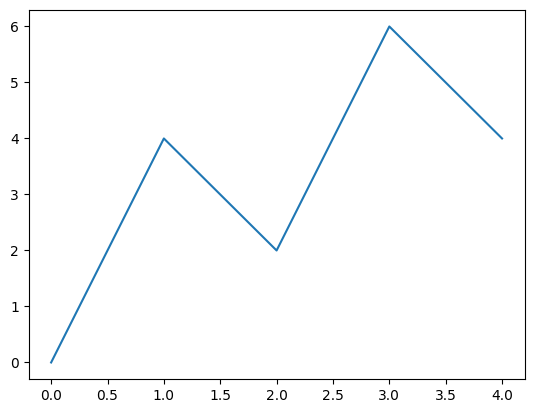
x = np.array([0,1,2,3,4])
y = np.array([0,4,2,6,4])
plt.plot(x,y)
[<matplotlib.lines.Line2D at 0x79200e81f750>]
Some formatting arguments include:
corcolor: line color (options:'k'or'black'for black,'red'for red, etc. – see this page for color options)lworlinewidth: line width (a number; the default is 1.5)lsorlinestyle: line style (options:'-', '--', '-.', ':')marker: optional marker style (options:'.', 'o', 'v', '^', '<', '>', 's', '*',etc.)msormarkersize: optional marker size (a number)
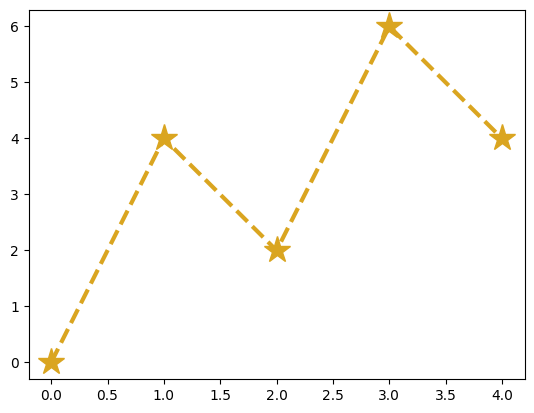
Try plotting x versus y again, except this time use a “goldenrod”-colored dashed line of width 2.5 with star-shaped markers of size 20:
# Write your code here:
plt.plot(x, y, color='goldenrod', lw=3, ls='--', marker='*', ms=20)
[<matplotlib.lines.Line2D at 0x791ff8b4e010>]
Some other options include changing the figure size by starting with a call to:
plt.figure(figsize=(WIDTH,HEIGHT))
Adding x-axis and y-axis labels and a title at the top:
plt.xlabel(STRING)
plt.ylabel(STRING)
plt.title(STRING)
Adding grid lines using:
plt.grid()
Or adding a plot legend by specifying the label argument in plt.plot() and adding using:
plt.legend()
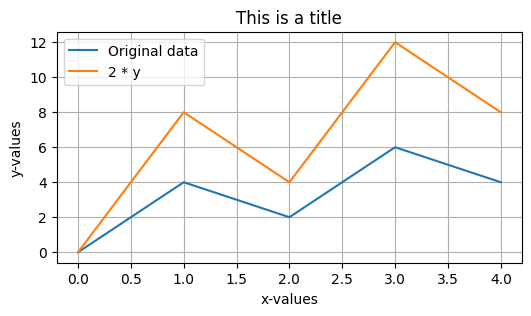
Check out these additional formatting options below:
plt.figure(figsize=(6,3))
plt.plot(x, y, label='Original data')
plt.plot(x, 2*y, label='2 * y') # y-values are multiplied by 2 here
plt.legend()
plt.grid()
plt.xlabel('x-values')
plt.ylabel('y-values')
plt.title('This is a title');
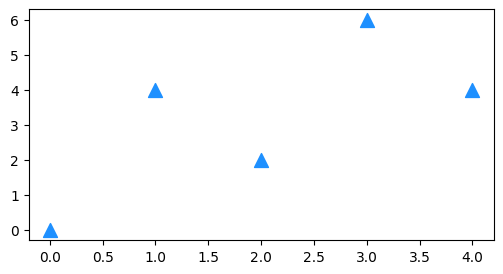
We can also create a scatter plot with just the points (no line). The function is similar to plt.plot():
plt.scatter(X, Y, s=SIZE, c=COLOR, marker=MARKER_STYLE, etc.)
plt.figure(figsize=(6,3))
plt.scatter(x, y, s=100, c='dodgerblue', marker='^');
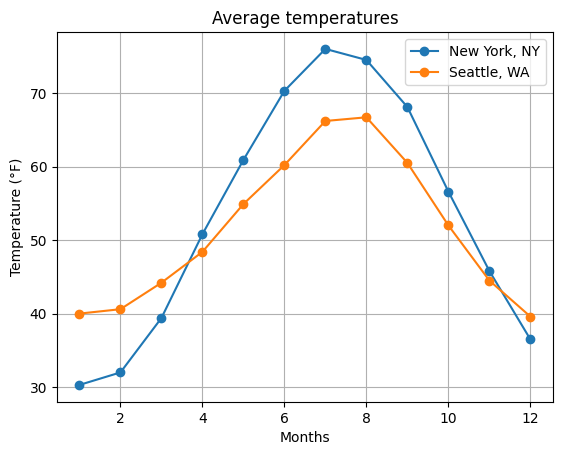
Let’s bring it all together! Below, try plotting the monthly temperatures in New York, NY and Seattle, WA. Use line plots with circle-shaped markers (or add scatter points separately). Include a legend and label the plot appropriately.
# Monthly temperatures in °F
temp = np.array([
[30.3,32.0,39.4,50.8,60.9,70.3,76.0,74.5,68.1,56.6,45.8,36.5], # New York
[40.0,40.6,44.2,48.4,54.9,60.2,66.2,66.7,60.5,52.0,44.5,39.6] # Seattle
])
# Write your code below:
months = np.arange(1,13)
plt.plot(months, temp[0,:], marker='o', label='New York, NY')
plt.plot(months, temp[1,:], marker='o', label='Seattle, WA')
plt.legend()
plt.xlabel('Months')
plt.ylabel('Temperature (°F)')
plt.title('Average temperatures')
plt.grid()
R/V Rachel Carson CTD depth profiles#
Image source: Emilio Mayorga, DINO SIP (SURP predecessor) 2024 cruise
First, let’s download two .csv data files from Google Drive here (we already used the two Rachel Carson files in the previous notebook!). Each file is a conductivity-temperature-depth (CTD) cast that was collected from the ship R/V Rachel Carson off of Carkeek Park near Seattle. There are 4 csv files on that folders; go ahead and save all 4 to your computer.
Next, we can upload the files to this Google Colab notebook. Click the sidebar folder icon on the left, then use the page-with-arrow icon
at the top to select the files and upload them.
Note that uploaded files will be deleted from Google Colab when you refresh this notebook!
We will specify each filepath using string variables:
Now, let’s plot the ocean CTD profiles measured by the R/V Rachel Carson. First we’ll read the two CTD csv files using pandas read_csv, as we did in the previous notebook.
Let’s remind ourselves of what the pandas DataFrame looks like:
filepath_1 = '/content/2023051001001_Carkeek.csv'
filepath_2 = '/content/2023051101001_Carkeek.csv'
data_1 = pd.read_csv(filepath_1, comment='#')
data_2 = pd.read_csv(filepath_2, comment='#')
# Note: in a notebook, we don't actually need the "display()" function
# to print out a variable (including a DataFrame) with nice formatting
data_1
| Unnamed: 0 | index | altM | CStarTr0 | c0mS/cm | density00 | depSM | latitude | longitude | flECO-AFL | ... | sbeox0Mg/L | sbeox0ML/L | ph | potemp090C | prDM | sal00 | t090C | scan | nbf | flag | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 0 | 3407 | 98.53 | 71.0825 | 31.662958 | 1021.7317 | 2.101 | 47.71418 | -122.40854 | 2.8127 | ... | 10.6450 | 7.4488 | 9.271 | 10.2155 | 2.119 | 28.3385 | 10.2157 | 3408 | 0 | 0.0 |
| 1 | 1 | 3408 | 98.53 | 71.0825 | 31.662061 | 1021.7317 | 2.005 | 47.71418 | -122.40854 | 2.8127 | ... | 10.6446 | 7.4484 | 9.271 | 10.2140 | 2.022 | 28.3388 | 10.2143 | 3409 | 0 | 0.0 |
| 2 | 2 | 3409 | 98.53 | 71.0825 | 31.661464 | 1021.7323 | 2.045 | 47.71418 | -122.40854 | 2.8127 | ... | 10.6443 | 7.4483 | 9.271 | 10.2129 | 2.062 | 28.3391 | 10.2131 | 3410 | 0 | 0.0 |
| 3 | 3 | 3410 | 98.53 | 71.0825 | 31.660448 | 1021.7323 | 2.005 | 47.71418 | -122.40854 | 2.8713 | ... | 10.6441 | 7.4481 | 9.271 | 10.2117 | 2.022 | 28.3390 | 10.2119 | 3411 | 0 | 0.0 |
| 4 | 4 | 3411 | 98.53 | 71.0825 | 31.658416 | 1021.7325 | 1.981 | 47.71418 | -122.40854 | 3.1057 | ... | 10.6443 | 7.4483 | 9.271 | 10.2093 | 1.998 | 28.3389 | 10.2095 | 3412 | 0 | 0.0 |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 8200 | 8200 | 11607 | 11.99 | 83.1087 | 31.920640 | 1024.1134 | 173.726 | 47.71316 | -122.40812 | 0.1753 | ... | 7.0198 | 4.9120 | 8.788 | 8.3719 | 175.266 | 30.0190 | 8.3887 | 11608 | 0 | 0.0 |
| 8201 | 8201 | 11608 | 11.99 | 83.1087 | 31.920640 | 1024.1135 | 173.726 | 47.71316 | -122.40812 | 0.1753 | ... | 7.0201 | 4.9123 | 8.788 | 8.3717 | 175.266 | 30.0191 | 8.3886 | 11609 | 0 | 0.0 |
| 8202 | 8202 | 11609 | 11.99 | 83.1087 | 31.920820 | 1024.1141 | 173.846 | 47.71316 | -122.40812 | 0.1753 | ... | 7.0204 | 4.9125 | 8.788 | 8.3718 | 175.387 | 30.0191 | 8.3887 | 11610 | 0 | 0.0 |
| 8203 | 8203 | 11610 | 11.99 | 83.1087 | 31.920579 | 1024.1129 | 173.613 | 47.71316 | -122.40812 | 0.1753 | ... | 7.0205 | 4.9125 | 8.783 | 8.3719 | 175.152 | 30.0190 | 8.3887 | 11611 | 0 | 0.0 |
| 8204 | 8204 | 11611 | 11.99 | 83.1087 | 31.920340 | 1024.1135 | 173.846 | 47.71316 | -122.40812 | 0.1753 | ... | 7.0209 | 4.9128 | 8.788 | 8.3720 | 175.387 | 30.0184 | 8.3889 | 11612 | 0 | 0.0 |
8205 rows × 21 columns
With the tools we have, we can make line plots of temperature vs. depth that include both CTD casts.
In the code below, we explicitly label the x- and y- axes.
plt.plot(data_1['t090C'], data_1['depSM'])
plt.plot(data_2['t090C'], data_2['depSM'])
plt.xlabel('Temperature (°C)')
plt.ylabel('Depth (m)')
Text(0, 0.5, 'Depth (m)')
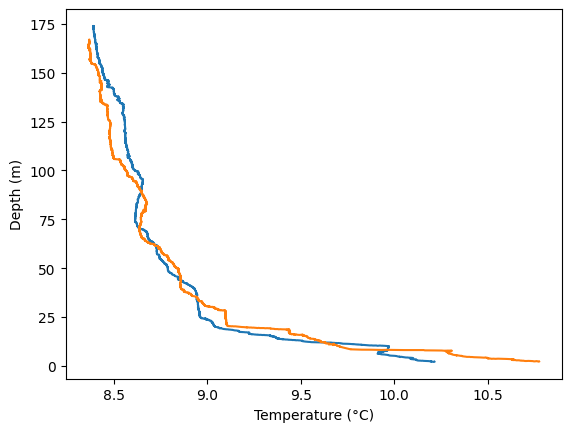
Hmm, it’d be more intuitive to have depth increasing from 0 at the top (the surface); and more useful to add a legend that clarifies which cast is which, a plot title and a grid.
# Temperature vs. depth profile
plt.plot(data_1['t090C'], data_1['depSM'], label='Cast #1 (5/10/23)')
plt.plot(data_2['t090C'], data_2['depSM'], label='Cast #2 (5/11/23)')
plt.xlabel('Temperature (°C)')
plt.ylabel('Depth (m)')
plt.title('R/V Carson CTD casts')
plt.legend()
plt.gca().invert_yaxis() # This reverses the y-axis. gca stands for "get current axes"
plt.grid()
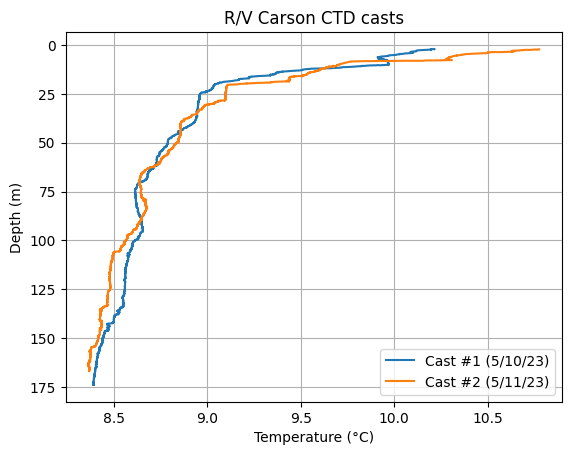
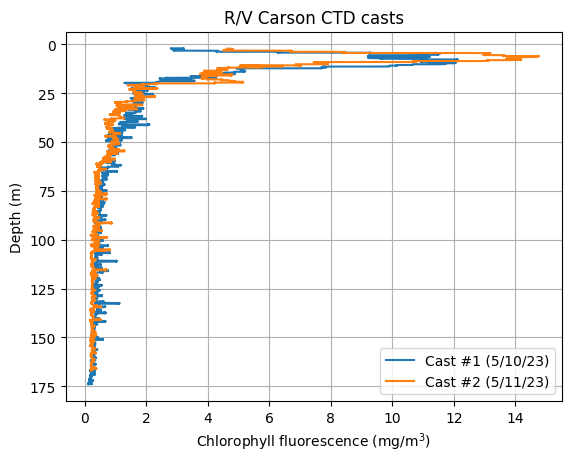
Can you try plotting another parameter vs. depth? Note: the file contains salinity (sal00), oxygen (sbeox0Mg/L), chlorophyll fluorescence (flECO-AFL), and pH (ph) data.
# Write your code here:
plt.figure()
plt.plot(data_1['flECO-AFL'], data_1['depSM'], label='Cast #1 (5/10/23)')
plt.plot(data_2['flECO-AFL'], data_2['depSM'], label='Cast #2 (5/11/23)')
plt.xlabel('Chlorophyll fluorescence (mg/m$^3$)')
plt.ylabel('Depth (m)')
plt.legend()
plt.gca().invert_yaxis() # This reverses the y-axis
plt.title('R/V Carson CTD casts')
plt.grid()
Part 2. Exploring ocean time series data from the Seattle Aquarium#
Seattle is located in King County. King County’s Department of Natural Resources & Parks maintains several ocean measurement stations in Puget Sound. These sensors monitor the water quality and ocean conditions.
One of these stations is at the Seattle Aquarium on the waterfront in downtown Seattle. The station consists of a mooring with two sensors. Sensor #1 is at a depth of 1 meter, and sensor #2 is at a depth of 10 m.
The mooring data can be obtained from King County here: https://green2.kingcounty.gov/marine-buoy/Data.aspx. However, the data requires a bit of processing before it can be loaded into Python. The data can also be conveniently visualized on the NANOOS Visualization System Data Explorer, here.
You can download the processed data file from Google Drive here. This CSV file, SeattleAquarium_7_2_2025_to_7_15_2025.csv, contains data measured every 15 minutes for the two weeks from July 2 to July 15, 2025.
Image source: MyEdmondsNews
The following call to pd.read_csv() will load the data file correctly.
The function arguments will ignore comments (comment='*'), set the header to the first non-commented row (header=0), set the index to the first column (index_col=0), interpret that column as datetimes (parse_dates=True), and specify the file input encoding (encoding='unicode_escape').
# Run this code to load the data
# When a function uses many arguments, it can be convenient for readability
# to specify each argument assignment in one line, like this:
aqua = pd.read_csv(
'/content/SeattleAquarium_7_2_2025_to_7_15_2025.csv',
comment='*',
header=0,
index_col=0,
parse_dates=True,
encoding='unicode_escape'
)
# .head(n) displays the first n records, where the default is n=5
# .tail(n) diplays the last n records.
aqua.head()
/tmp/ipython-input-2993415281.py:4: UserWarning: Could not infer format, so each element will be parsed individually, falling back to `dateutil`. To ensure parsing is consistent and as-expected, please specify a format.
aqua = pd.read_csv(
| 1_Depth_m | Qual_1_Depth | 2_Depth_m | Qual_2_Depth | 1_Water_Temperature_degC | Qual_1_Water_Temperature | 2_Water_Temperature_degC | Qual_2_Water_Temperature | 1_Salinity_PSU | Qual_1_Salinity | ... | Qual_2_Sonde_pH | 1_Density_kg/m^3 | Qual_1_Water_Density | 2_Density_kg/m^3 | Qual_2_Water_Density | 1_Sonde_Batt_V | 2_Sonde_Batt_V | Logger_Batt_V | 1_Sonde_ID | 2_Sonde_ID | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Date | |||||||||||||||||||||
| 2025-07-02 00:00:00 | 0.852 | 210 | 9.799 | 210 | 13.415 | 210 | 11.443 | 210 | 29.021 | 210 | ... | 210 | 1021.68965 | 210 | 1022.65046 | 210 | 11.9 | 13.8 | 13.1 | NaN | NaN |
| 2025-07-02 00:15:00 | 0.845 | 210 | 9.769 | 210 | 13.636 | 210 | 11.456 | 210 | 28.910 | 210 | ... | 210 | 1021.56106 | 210 | 1022.63347 | 210 | 11.9 | 13.8 | 13.1 | NaN | NaN |
| 2025-07-02 00:30:00 | 0.844 | 210 | 9.791 | 210 | 13.356 | 210 | 11.447 | 210 | 29.042 | 210 | ... | 210 | 1021.71730 | 210 | 1022.63654 | 210 | 11.9 | 13.8 | 13.1 | NaN | NaN |
| 2025-07-02 00:45:00 | 0.865 | 210 | 9.811 | 210 | 13.274 | 210 | 11.422 | 210 | 29.080 | 210 | ... | 210 | 1021.76240 | 210 | 1022.65867 | 210 | 11.9 | 13.8 | 13.1 | NaN | NaN |
| 2025-07-02 01:00:00 | 0.846 | 210 | 9.823 | 210 | 13.298 | 210 | 11.390 | 210 | 29.061 | 210 | ... | 210 | 1021.74305 | 210 | 1022.67361 | 210 | 11.9 | 13.8 | 13.1 | NaN | NaN |
5 rows × 39 columns
Since Pandas won’t display all the column names (there are too many!), we can use the .columns attribute to see them:
# Note: we don't need "print()" or "display()" ;)
aqua.columns
Index(['1_Depth_m', 'Qual_1_Depth', '2_Depth_m', 'Qual_2_Depth',
'1_Water_Temperature_degC', 'Qual_1_Water_Temperature',
'2_Water_Temperature_degC', 'Qual_2_Water_Temperature',
'1_Salinity_PSU', 'Qual_1_Salinity', '2_Salinity_PSU',
'Qual_2_Salinity', '1_Dissolved_Oxygen_%Sat', '1_Dissolved_Oxygen_mg/L',
'Qual_1_DO', '2_Dissolved_Oxygen_%Sat', '2_Dissolved_Oxygen_mg/L',
'Qual_2_DO', '1_Chlorophyll_Fluorescence_ug/L',
'Qual_1_Chlorophyll_Fluorescence', '2_Chlorophyll_Fluorescence_ug/L',
'Qual_2_Chlorophyll_Fluorescence', '1_Turbidity_NTU',
'Qual_1_Turbidity', '2_Turbidity_NTU', 'Qual_2_Turbidity', '1_Sonde_pH',
'Qual_1_Sonde_pH', '2_Sonde_pH', 'Qual_2_Sonde_pH', '1_Density_kg/m^3',
'Qual_1_Water_Density', '2_Density_kg/m^3', 'Qual_2_Water_Density',
'1_Sonde_Batt_V', '2_Sonde_Batt_V', 'Logger_Batt_V', '1_Sonde_ID',
'2_Sonde_ID'],
dtype='object')
We can also call .info() to get even more information about the DataFrame in a nicely formatted presentation:
aqua.info()
<class 'pandas.core.frame.DataFrame'>
DatetimeIndex: 1298 entries, 2025-07-02 00:00:00 to 2025-07-15 13:15:00
Data columns (total 39 columns):
# Column Non-Null Count Dtype
--- ------ -------------- -----
0 1_Depth_m 1298 non-null float64
1 Qual_1_Depth 1298 non-null int64
2 2_Depth_m 1298 non-null float64
3 Qual_2_Depth 1298 non-null int64
4 1_Water_Temperature_degC 1298 non-null float64
5 Qual_1_Water_Temperature 1298 non-null int64
6 2_Water_Temperature_degC 1298 non-null float64
7 Qual_2_Water_Temperature 1298 non-null int64
8 1_Salinity_PSU 1298 non-null float64
9 Qual_1_Salinity 1298 non-null int64
10 2_Salinity_PSU 1298 non-null float64
11 Qual_2_Salinity 1298 non-null int64
12 1_Dissolved_Oxygen_%Sat 1298 non-null float64
13 1_Dissolved_Oxygen_mg/L 1298 non-null float64
14 Qual_1_DO 1298 non-null int64
15 2_Dissolved_Oxygen_%Sat 1298 non-null float64
16 2_Dissolved_Oxygen_mg/L 1298 non-null float64
17 Qual_2_DO 1298 non-null int64
18 1_Chlorophyll_Fluorescence_ug/L 1298 non-null float64
19 Qual_1_Chlorophyll_Fluorescence 1298 non-null int64
20 2_Chlorophyll_Fluorescence_ug/L 1298 non-null float64
21 Qual_2_Chlorophyll_Fluorescence 1298 non-null int64
22 1_Turbidity_NTU 1298 non-null float64
23 Qual_1_Turbidity 1298 non-null int64
24 2_Turbidity_NTU 1298 non-null float64
25 Qual_2_Turbidity 1298 non-null int64
26 1_Sonde_pH 1298 non-null float64
27 Qual_1_Sonde_pH 1298 non-null int64
28 2_Sonde_pH 1298 non-null float64
29 Qual_2_Sonde_pH 1298 non-null int64
30 1_Density_kg/m^3 1297 non-null float64
31 Qual_1_Water_Density 1298 non-null int64
32 2_Density_kg/m^3 1297 non-null float64
33 Qual_2_Water_Density 1298 non-null int64
34 1_Sonde_Batt_V 1298 non-null float64
35 2_Sonde_Batt_V 1298 non-null float64
36 Logger_Batt_V 1298 non-null float64
37 1_Sonde_ID 0 non-null float64
38 2_Sonde_ID 0 non-null float64
dtypes: float64(23), int64(16)
memory usage: 405.6 KB
We learned how to create X-Y line and scatter plots using plt.plot() above. However, Pandas offers us a shortcut.
You can call .plot() on a Pandas Series to generate a line plot. The function arguments include many of those you learned for plt.plot(). They can be found in the online documentation: https://pandas.pydata.org/docs/reference/api/pandas.Series.plot.html.
Pandas .plot() offers some advantages. It’s simpler to use, and as you’ll see it auto-generates axis labels and legends, as needed.
# Plot temperature from upper (1-meter) and lower (10-meter) sensors
aqua['1_Water_Temperature_degC'].plot() # 1-meter
aqua['2_Water_Temperature_degC'].plot() # 10-meter
plt.legend()
<matplotlib.legend.Legend at 0x791ff85e8450>
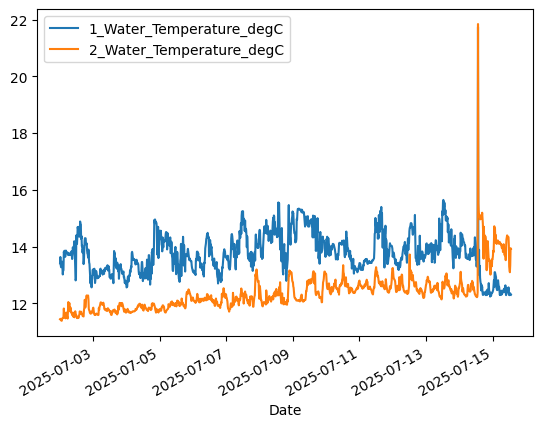
Note the spike in the temperature data on July 14, and then the apparent switch of the Sensor 1 and Sensor 2 data streams. What do you think happened at the Aquarium?
We’ll learn how to use boolean indexing to mask out bad data later. But for now, let’s change the y-axis limits to zoom in on the good data.
# Plot temperature from upper (1-meter) sensor
aqua['1_Water_Temperature_degC'].plot()
aqua['2_Water_Temperature_degC'].plot()
plt.ylim([11.0,16.0]) # Set y-axis limits to ignore data spike
plt.legend()
<matplotlib.legend.Legend at 0x791ff85a4c50>
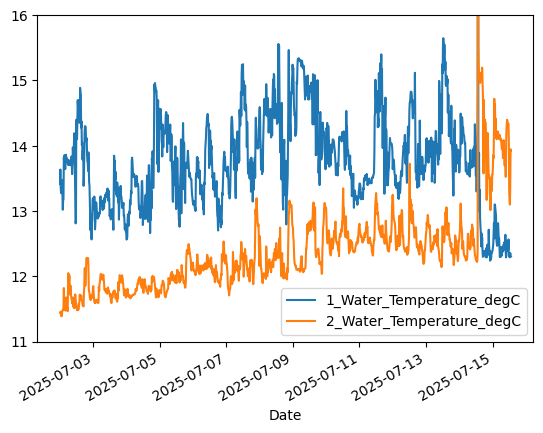
Using timeseries capabilities#
We can take advantage of built-in datetime functionality in Pandas to examine data at aggregated time intervals, like daily; and to examine diurnal variability (within each day).
First let’s examine the DataFrame index:
# The DataFrame index has a numpy "datetime64[ns]" data type
aqua.index
DatetimeIndex(['2025-07-02 00:00:00', '2025-07-02 00:15:00',
'2025-07-02 00:30:00', '2025-07-02 00:45:00',
'2025-07-02 01:00:00', '2025-07-02 01:15:00',
'2025-07-02 01:30:00', '2025-07-02 01:45:00',
'2025-07-02 02:00:00', '2025-07-02 02:15:00',
...
'2025-07-15 11:00:00', '2025-07-15 11:15:00',
'2025-07-15 11:30:00', '2025-07-15 11:45:00',
'2025-07-15 12:00:00', '2025-07-15 12:15:00',
'2025-07-15 12:30:00', '2025-07-15 12:45:00',
'2025-07-15 13:00:00', '2025-07-15 13:15:00'],
dtype='datetime64[ns]', name='Date', length=1298, freq=None)
# Extract the date from each datetime value in the index using the .date property
aqua.index.date
array([datetime.date(2025, 7, 2), datetime.date(2025, 7, 2),
datetime.date(2025, 7, 2), ..., datetime.date(2025, 7, 15),
datetime.date(2025, 7, 15), datetime.date(2025, 7, 15)],
dtype=object)
We can use .resample() to resample the DataFrame over a specific time resolution (“1D” = 1 day), returning the minimum (.min()) and maximum (.max()) values at the resampled resolution:
aqua_dailymin = aqua.resample("1D", origin='start_day').min()
aqua_dailymax = aqua.resample("1D", origin='start_day').max()
We now have DataFrames with just one record (row) per day!
aqua_dailymin
| 1_Depth_m | Qual_1_Depth | 2_Depth_m | Qual_2_Depth | 1_Water_Temperature_degC | Qual_1_Water_Temperature | 2_Water_Temperature_degC | Qual_2_Water_Temperature | 1_Salinity_PSU | Qual_1_Salinity | ... | Qual_2_Sonde_pH | 1_Density_kg/m^3 | Qual_1_Water_Density | 2_Density_kg/m^3 | Qual_2_Water_Density | 1_Sonde_Batt_V | 2_Sonde_Batt_V | Logger_Batt_V | 1_Sonde_ID | 2_Sonde_ID | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Date | |||||||||||||||||||||
| 2025-07-02 | 0.804 | 210 | 9.769 | 210 | 12.565 | 210 | 11.388 | 210 | 27.520 | 210 | ... | 210 | 1020.44613 | 210 | 1022.25050 | 210 | 11.7 | 13.8 | 13.0 | NaN | NaN |
| 2025-07-03 | 0.862 | 210 | 9.816 | 210 | 12.683 | 210 | 11.590 | 210 | 28.036 | 210 | ... | 210 | 1020.95768 | 210 | 1022.39893 | 210 | 11.7 | 13.8 | 13.0 | NaN | NaN |
| 2025-07-04 | 0.882 | 210 | 9.839 | 210 | 12.563 | 210 | 11.662 | 210 | 27.327 | 210 | ... | 210 | 1020.31357 | 210 | 1022.34168 | 210 | 11.6 | 13.8 | 13.0 | NaN | NaN |
| 2025-07-05 | 0.976 | 210 | 9.925 | 210 | 12.759 | 210 | 11.680 | 210 | 24.558 | 210 | ... | 210 | 1018.04794 | 210 | 1022.19333 | 210 | 11.3 | 13.8 | 13.0 | NaN | NaN |
| 2025-07-06 | 0.912 | 210 | 9.872 | 210 | 12.702 | 210 | 11.840 | 210 | 27.261 | 210 | ... | 210 | 1020.12523 | 210 | 1022.28316 | 210 | 11.5 | 13.8 | 13.0 | NaN | NaN |
| 2025-07-07 | 0.955 | 210 | 9.889 | 210 | 13.146 | 210 | 11.709 | 210 | 27.778 | 210 | ... | 210 | 1020.39621 | 210 | 1022.06330 | 210 | 11.5 | 13.8 | 13.0 | NaN | NaN |
| 2025-07-08 | 0.913 | 210 | 9.869 | 210 | 12.795 | 210 | 11.896 | 210 | 27.748 | 210 | ... | 210 | 1020.40477 | 210 | 1021.99987 | 210 | 11.5 | 13.8 | 13.0 | NaN | NaN |
| 2025-07-09 | 0.926 | 210 | 9.902 | 210 | 13.398 | 210 | 12.038 | 210 | 27.377 | 210 | ... | 210 | 1020.06360 | 210 | 1022.05351 | 210 | 11.5 | 13.8 | 13.0 | NaN | NaN |
| 2025-07-10 | 0.923 | 210 | 9.878 | 210 | 13.050 | 210 | 12.309 | 210 | 27.690 | 210 | ... | 210 | 1020.53203 | 210 | 1021.83855 | 210 | 11.2 | 13.8 | 13.0 | NaN | NaN |
| 2025-07-11 | 0.840 | 210 | 9.804 | 210 | 13.180 | 210 | 12.318 | 210 | 27.572 | 210 | ... | 210 | 1020.30141 | 210 | 1021.94333 | 210 | 11.5 | 13.8 | 13.0 | NaN | NaN |
| 2025-07-12 | 0.923 | 210 | 9.875 | 210 | 13.189 | 210 | 12.191 | 210 | 28.267 | 210 | ... | 210 | 1020.82972 | 210 | 1021.82511 | 210 | 11.2 | 13.8 | 13.0 | NaN | NaN |
| 2025-07-13 | 0.858 | 210 | 9.804 | 210 | 13.237 | 210 | 12.141 | 210 | 27.482 | 210 | ... | 210 | 1020.05341 | 210 | 1022.25472 | 210 | 11.5 | 13.8 | 13.0 | NaN | NaN |
| 2025-07-14 | -0.010 | 210 | -0.011 | 210 | 12.240 | 210 | 12.220 | 210 | 0.013 | 210 | ... | 210 | 1021.38435 | 210 | 1020.60829 | 210 | 11.5 | 13.3 | 13.0 | NaN | NaN |
| 2025-07-15 | 9.642 | 330 | 0.716 | 330 | 12.291 | 210 | 13.099 | 210 | 29.540 | 210 | ... | 210 | 1022.19247 | 330 | 1020.99987 | 330 | 12.0 | 13.4 | 13.0 | NaN | NaN |
14 rows × 39 columns
Look at the large changes in depth for the two sensors, 1_Depth_m and 2_Depth_m, starting on July 14!
Use the resampled daily DataFrames to plot the 1-meter daily minimum and maximum on top of the raw data at the original 15-minute resolution.
aqua['1_Water_Temperature_degC'].plot(label="Raw data")
aqua_dailymin['1_Water_Temperature_degC'].plot(label="Daily min")
aqua_dailymax['1_Water_Temperature_degC'].plot(label="Daily max")
plt.legend()
plt.ylabel("1_Water_Temperature_degC")
plt.title("Water temperature at 1-meter depth")
plt.grid()
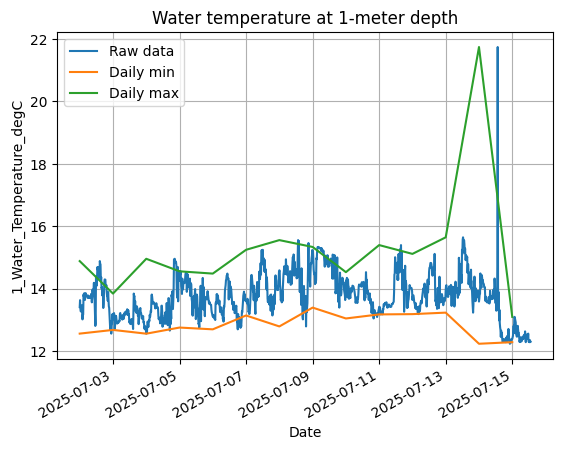
Now let’s focus on how temperature varies throughout the day (“diurnally”). First, use the DataFrame index to create an array of time values as fractional hours. For example, 12:30 is 12.5. We use the .hour and .minute properties:
hours_minutes = aqua.index.hour + aqua.index.minute / 60
hours_minutes
Index([ 0.0, 0.25, 0.5, 0.75, 1.0, 1.25, 1.5, 1.75, 2.0, 2.25,
...
11.0, 11.25, 11.5, 11.75, 12.0, 12.25, 12.5, 12.75, 13.0, 13.25],
dtype='float64', name='Date', length=1298)
# Plot temperature as a scatter plot
plt.scatter(
hours_minutes, aqua['2_Water_Temperature_degC'], label='10 meter',
s=30, c='orange', marker='^', alpha=0.2
)
plt.scatter(
hours_minutes, aqua['1_Water_Temperature_degC'], label='1 meter',
s=5, c='blue', marker='o', alpha=0.2
)
plt.ylim([11.0,16.0]) # Set y-axis limits to ignore data spike
plt.xlabel("Hour (0-24)")
plt.ylabel("Water Temperature (oC)")
plt.legend()
plt.title(
f"Diurnal temperature variability, {aqua.index.min():%B %d} to {aqua.index.max():%B %d}"
);
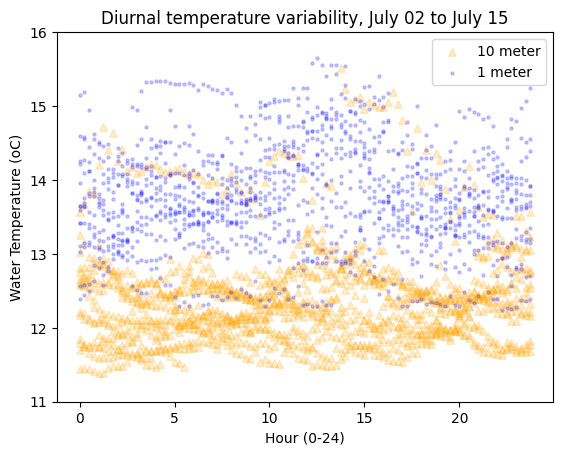
In the title, we used datetime formatting codes with f string formatting. See https://strftime.org and https://www.strfti.me. Here’s a simpler example:
f"{aqua.index.min():%B %d}"
'July 02'
The slice() function can be convenient for generating indices over an interval. The syntax is:
slice(start, stop, step=None)
For example, we can apply a slice on the timeseries DataFrame index to plot 1-meter temperature from the start of the timeseries to July 9. Notice that we use None to start at the beginning of the time series and can define a date as a string!
aqua['1_Water_Temperature_degC'].loc[slice(None, '2025-07-09')].plot(
label="Raw data", figsize=(8,3)
);
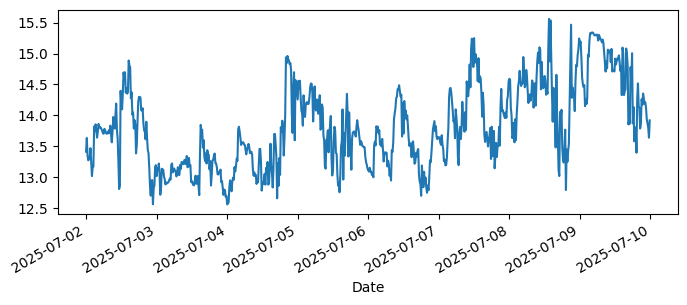
Try exploring the data#
Can you answer some of the following questions by making plots and using the functions you already know?
What was the warmest ocean temperature seen in this data? (Feel free to slice the data to ignore periods of seemingly incorrect measurements.)
On average, how much colder is the deep (10-meter) sensor than the shallow (1-meter) sensor? (Feel free to slice the data to ignore periods of seemingly incorrect measurements.)
What day likely had a significant rain event? (Hint: rain is fresh water, and the ocean is salty.)
What direction is phytoplankton growth trending in over this data period? (Hint: chlorophyll concentration is a measure of how much phytoplankton are in seawater.)
What is the relationship between near-surface ocean temperature and phytoplankton?
Can you plot the mean daily oxygen concentration at 1- and 10-meter depths (the two
Dissolved_Oxygen_mg/Lcolumns)?
# Warmest ocean temperature
# Note: we are using .loc[slice()] to slice the data in time
# to ignore bad data from July 13 onwards
print(aqua['1_Water_Temperature_degC'].loc[slice(None,'2025-07-13')].max())
print(aqua['2_Water_Temperature_degC'].loc[slice(None,'2025-07-13')].max())
15.648
13.724
# Average temperature difference between shallow and deep sensors
# Note: we are using .loc[slice()] to slice the data in time
# to ignore bad data from July 13 onwards
print(aqua['1_Water_Temperature_degC'].loc[slice(None,'2025-07-13')].mean() -
aqua['2_Water_Temperature_degC'].loc[slice(None,'2025-07-13')].mean())
1.6307517421602782
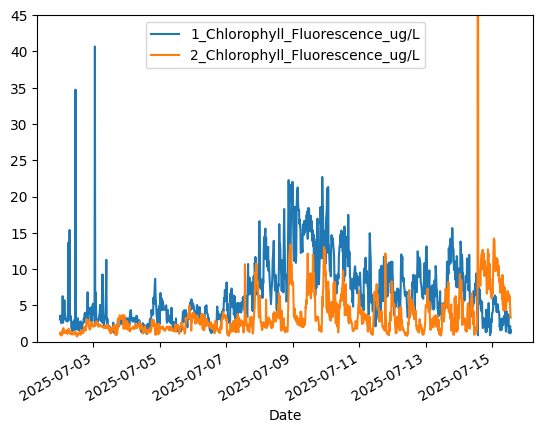
# Phytoplankton growth trend appears to be initially upwards, then downwards after July 9
aqua['1_Chlorophyll_Fluorescence_ug/L'].plot()
aqua['2_Chlorophyll_Fluorescence_ug/L'].plot()
plt.ylim([0.0,45.0]) # Crop y-limits due to data spike
plt.legend()
<matplotlib.legend.Legend at 0x791ff8301290>
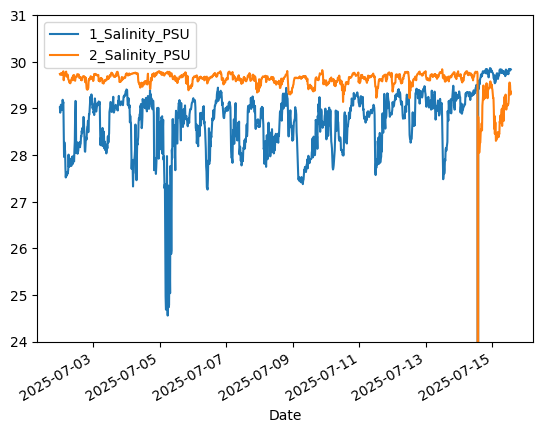
# Ocean salinity reveals a freshening signal on 7/5/25, likely from rain
aqua['1_Salinity_PSU'].plot()
aqua['2_Salinity_PSU'].plot()
plt.ylim([24,31]) # Crop y-limits due to data spike
plt.legend()
<matplotlib.legend.Legend at 0x791ff820f210>
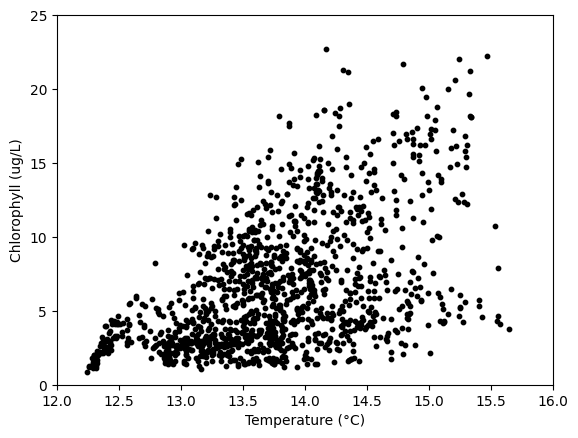
# Relationship between temperature and chlorophyll
# (The amount of phytoplankton seems to increase with warmer
# near-surface temperatures!)
plt.scatter(aqua['1_Water_Temperature_degC'],
aqua['1_Chlorophyll_Fluorescence_ug/L'],
c='k',s=10)
plt.xlim([12.0,16.0]) # Crop axis limits due to incorrect data
plt.ylim([0.0,25.0])
plt.xlabel('Temperature (°C)')
plt.ylabel('Chlorophyll (ug/L)')
Text(0, 0.5, 'Chlorophyll (ug/L)')
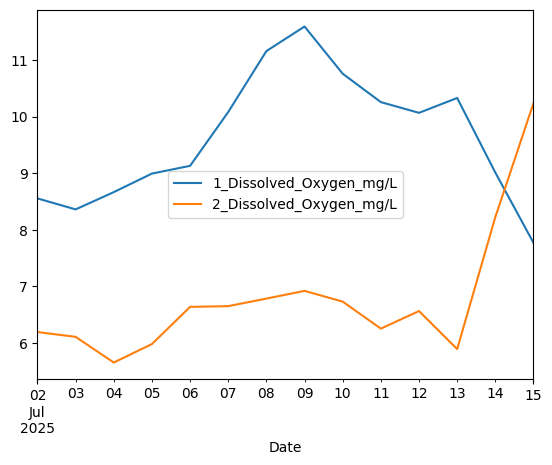
# Mean daily oxygen at 1 and 10 meters
aqua_dailymean = aqua.resample("1D", origin='start_day').mean()
aqua_dailymean[['1_Dissolved_Oxygen_mg/L', '2_Dissolved_Oxygen_mg/L']].plot()
<Axes: xlabel='Date'>
Part 3: Logical operations#
Often, we will want to compare two numbers or variables. We do this using the following logical operations:
==: equal!=: not equal>: greater than>=: greater than or equal to<: less than<=: less than or equal toandor&: are both booleans true?oror|: is either boolean true?notor~: reverse the boolean (True -> False, False -> True)in: is a membernot in: is not a member
Each logical operation evaluates to (returns) a boolean — True or False. Consider the following examples:
3 == 3
True
3 == 3.0 # integers can be compared to floating-point numbers
True
not 3 == 3
False
3 == 5
False
3 != 5
True
3 > 5
False
5 <= 5
True
(3 != 5) or (3 > 5)
True
(3 != 5) and (3 > 5)
False
Applying a logical comparison to a NumPy array gives a boolean array!
x = np.array([1,2,3,4,5,6])
print(x < 4)
print(x <= 4)
[ True True True False False False]
[ True True True True False False]
# Note: "not" can't be applied to an entire boolean array.
# Instead, we have to use "~":
print(~np.array([True, False, True]))
[False True False]
Note that membership tests work on lists, arrays, and strings:
print(3 in x) # this is asking: "is 3 in x?"
True
print(7 in x)
False
print(3 not in x) # this is asking: "is 3 not in x?"
False
print('o w' in 'hello world')
True
print('World' in 'hello world') # note that string membership is case-sensitive
False
Heads up: this next skill is super powerful. We saw above that applying a logical comparison to an array of numbers gives us a boolean array.
We can use boolean arrays as “masks” to select certain elements of an array. This is called boolean indexing.
Here are a few ways to use it:
# Here are the pH values from last week's lesson:
pH_measurements = np.array([7.84, 7.91, 8.05, np.nan, 7.96, 8.03])
print('pH measurements:', pH_measurements)
# Remember that we can test for missing data (np.NaN values) using np.isnan():
print('Result of np.isnan():', np.isnan(pH_measurements))
# The resulting boolean array can be used to extract only the valid data:
print('Array after removing missing data:', pH_measurements[~np.isnan(pH_measurements)])
pH measurements: [7.84 7.91 8.05 nan 7.96 8.03]
Result of np.isnan(): [False False False True False False]
Array after removing missing data: [7.84 7.91 8.05 7.96 8.03]
# Let's revisit the Seattle temperatures from earlier:
seattle_temps = np.array([40.0,40.6,44.2,48.4,54.9,60.2,66.2,66.7,60.5,52.0,44.5,39.6])
# Applying a logical comparison creates a boolean array, or "mask":
print(seattle_temps > 60)
[False False False False False True True True True False False False]
# Now let's use the mask to retrieve only the elements where the mask is True:
seattle_temps[seattle_temps > 60]
# Note: this only works when the mask is the same length as the array!
array([60.2, 66.2, 66.7, 60.5])
# The boolean indexing gives the same result as specifying the actual array indices:
seattle_temps[[5,6,7,8]]
array([60.2, 66.2, 66.7, 60.5])
We can use boolean indexing to handle the data outlier in the Seattle Aquarium time series! Let’s apply the threshold of 16°C we used earlier to set values as “missing”, np.nan. To do this, we’ll use the extended .loc[] syntax to assign a value to a column based on a criteria:
dataframe.loc[BOOLEAN_ARRAY, COLUMN_NAME] = NEW_VALUE
aqua.loc[aqua['1_Water_Temperature_degC'] > 16, '1_Water_Temperature_degC'] = np.nan
aqua.loc[aqua['2_Water_Temperature_degC'] > 16, '2_Water_Temperature_degC'] = np.nan
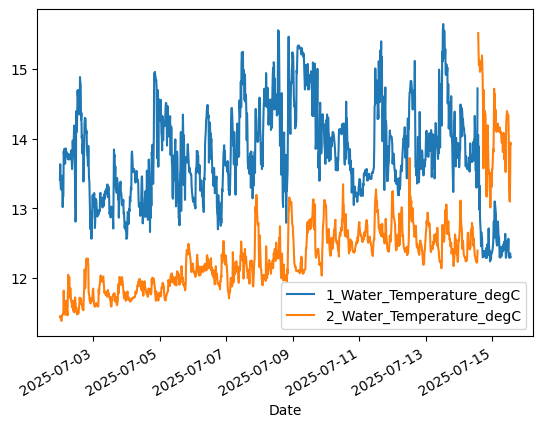
Now redo the plot we did earlier. No need to clip the y axis range anymore! But the data still seem to switchover between the two sensors …
aqua['1_Water_Temperature_degC'].plot() # 1-meter
aqua['2_Water_Temperature_degC'].plot() # 10-meter
plt.legend()
<matplotlib.legend.Legend at 0x791ff8314dd0>
How many months of the year is Seattle 40°F or colder? Try using boolean indexing and a function that you’ve learned to calculate and print the answer:
# Write your code here:
len(seattle_temps[seattle_temps > 40])
10

